University of Virginia |
CS Department at UVA |
Swarm programming @ CS @ UVA
Getting Started
This document contains the following sections
- Features
- Installation
- Running Cell Programs
- Writing Cell programs
- Writing models for CellSim
Conventions: Anything in [] is optional, Anything in <> is obligatory,
Anything within "" must be typed as is
- An architecture that allows dynamically loading a simulation using user-defined models for
- Cell program
- Cell Model
- System Model
- A sim / config file will load these into a
simulation object by reading a file with contents
system=sysmodels.simplebiology.SimpleBiology
cell=cellmodels.simplecell.SimpleCell
parser=parsers.CPA.CPAParser
- Better UI to allow simultaneous viewing of
several simulations (at different stages)
- Arcitecure allows easy specification of diffferent statistics
- Extract the cellsim.zip file to C:\cellsim\. If you install to a different path be sure to edit the init_env file present in the cellsim main directory to reflect your new path.
- Install J2SDK from http://java.sun.com/downloads/ [Ver 1.4.1]
- Install Java3D also from http://java.sun.com/downloads/
filename should be java3d-1_3-windows-i586-opengl-sdk.exe for OpenGL version for the i586
- Install JavaCC (2.1) from http://www.webgain.com/download/javacc/details.html
Note: This is not needed for running CellSim
All 3 are also available at http://www.cs.virginia.edu/~gsg5j/download/
The applet can be viewed once (1) & (2) are installed. The applet is available at
http://www.cs.virginia.edu/~gsg5j/cellsim/
The stand-alone version of CellSim is available at
http://swarm.cs.virginia.edu/cellsim/download/cellsim.zip
cd CellSim
init_env
java gui.GUI
Select File->New and then Simulation->Step. Using the menu File->New, you can run as many simulations as you like in seperate internal windows within the main window
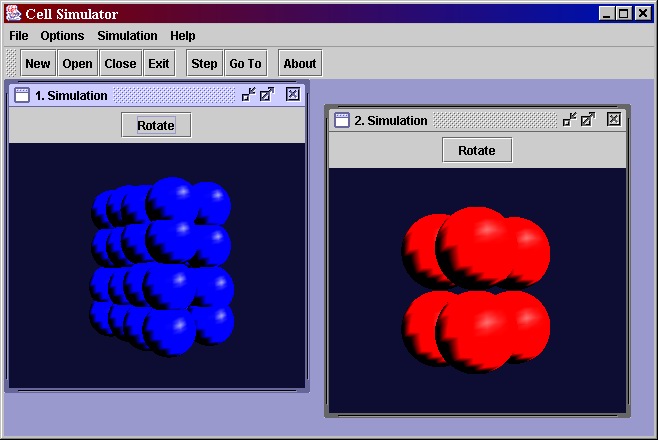
- It is better to draw a diagram of the cell state transitions as this will give a better idea of the control flow.
- Check the BNF for the CPA Grammar here.
- You can write your own cell program grammars by adding modules to the parsers directory. The next section describes how this can be done.
- The abstract classes SystemModel, Cell and Parser reside in the core/ subdirectory and form the heart of the Cell Simulator. These must be subclassed to form different models for the system, cell and the parser
- The class Simulation dynamically loads the models specified in the SIM file for the SystemModel, Cell and Parser
- To create your own system model, I suggest that you begin by creating a copy of the subdirectory sysmodels/simplebiology. Store it as sysmodels/MySystemModel. Rename the class-names and the package-name in sysmodels/MySystemModel/SimpleBiology.java suitably. The heart of the model is the function
public Vector provideService(Vector args)
It is basically a function called by the Cell to cause system-wide effects like chemical diffusion, cell diffusion etc
- You can similarly create a new Cell model by starting with cellmodels/SimpleCell/SimpleCell.java. Again begin by modifying the provideService() function
- Be sure to create a sim file similar to simple.sim provided in the main directory of the CellSim distribution. Also make sure that the classes for your new models are reacheable through the CLASSPATH. As long as you keep all cell models in the cellmodels/ folder, all system models in the sysmodels/ folder and all parsers in the parsers/ folder and name the packages using the pathname from the base directory, this should not be a problem